Georg Thimm, Elvira Gawlitta-Gorka, Gabriele Sorg, Horst Flotow
Hit Discovery Constance GmbH, Constance, Germany
www.hit-discovery.com
Introduction
In vivo, most cells interact with neighboring cells and extracellular matrix components to establish a unique three-dimensional (3D) organization that is critical for normal cell and tissue function. Cells grown in conventional two-dimensional (2D) monolayer cultures lose many of these tissue-specific properties. A number of 3D cell culture models have been developed, including organotypic (tissue) explants, spheroid cultures (hanging drop), and 3D cell culture in a variety of scaffolds (e.g. hydrogels, alginate sponges, spun fibers and others).
Although these 3D culture models are biochemically and physiologically more similar to in vivo tissues, there are still a number of technical challenges to overcome before they can be implemented in routine drug discovery processes such as cytotoxicity testing. In particular, 3D models tend to be complex and thus difficult to establish and maintain. The complexity and technical challenges limit the throughput and also make it difficult to compare results between different laboratories (lack of validation of assay robustness and reproducibility).
Here we have started addressing some of these challenges using GrowDex® hydrogel as a 3D cell culture matrix for growing liver-derived HepG2 cells in 384 well microtitre plates.
Materials and methods
- The 1.5% stock concentration of GrowDex (Cat no. 100 103 005, UPM) was diluted with assay medium to obtain a working concentration of 0.5% GrowDex. 250,000 HepG2 cells (Cat no. 85011430, Sigma) cells in 100 µl were added to 0.9 ml of the GrowDex working solution and mixed gently.
- 30 µl of the cell-containing mixture was dispensed into the wells of the 384 well assay plate (Cat no. 781091, Greiner) and left for 30 minutes at 37°C, then each well was overlaid with 50 µl assay medium.
- After an overnight incubation at 37°C, 40 µl of the overlaid (supernatant) medium was removed using a CyBiWell 384 well dispenser (without touching the GrowDex layer). 40 µl of test compounds (or appropriate DMSO controls) diluted in assay medium were added per well. The plates were then incubated at 37°C for 4 days.
- After 4 days, 40 µl of the supernatant was removed from all wells and the cells fixed using Prefer (Anatech Ltd.) according to the manufacturer’s instructions and the nuclei stained with DRAQ5 (Cat no. 62251, ThermoFischer). Fluorescent images were taken using the OPERA confocal automated microscope (Perkin Elmer; Excitation at 635 nm/Emission at 690 nm).
- Automated image analysis to determine the nuclear count (per field and per well) as a measure of cell viability was carried out using CellProfiler.
- The pilot screen used the above method, acquiring 4 slices (Z-stacks) of 4 fields in each well and determining the cell count in the resultant images in CellProfiler.
Results
Following some optimization, the protocols described in Materials and Methods were used to establish the assay in 384 well microtitre plates. The results shown below demonstrate a dose-dependent effect of a chemotherapeutic drug camptothecin (Cat no. C9911, Sigma) on the cell number. This can be seen clearly in the images (Fig. 1) with an IC50 value being quantified as 29nM (Fig. 2).
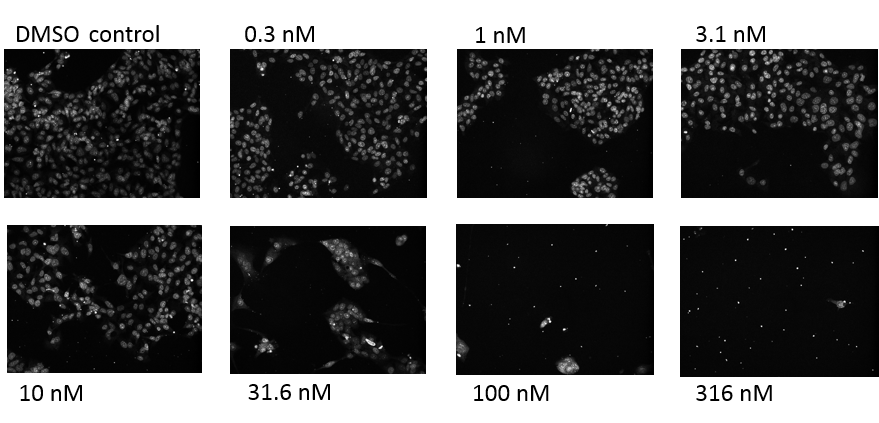
Figure 1. Nuclear staining of HepG2 cells in 3D cell culture with DRAQ5. Cells growing in the 384 well plates as described were treated for 4 days with concentrations of camptothecin ranging from 0.3 nM to 316 nM. The cells were fixed and stained, and images acquired using the OPERA system.
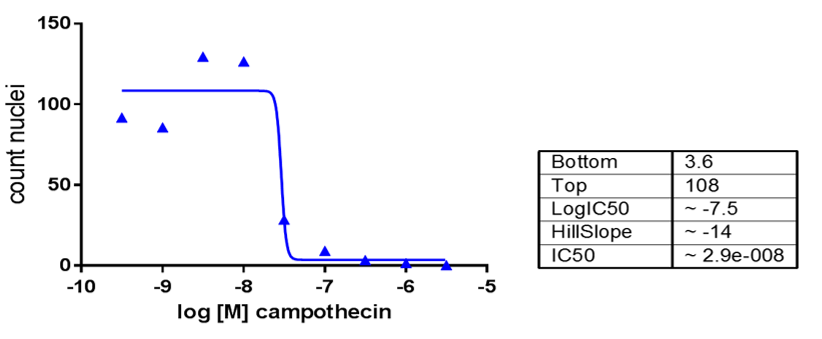
Figure 2. Dose dependent cytotoxic activity of camptothecin in HepG2 cells. The cell count (number of nuclei in the confocal images) was quantified in the images acquired using CellProfiler. The IC50 was calculated with GraphPad using the Hill equation.
A pilot screen against a small 1600 compound library at 20 µM compound concentration was run to demonstrate the utility of the assay. The pilot screen results are shown in Fig. 3 and Fig. 4.
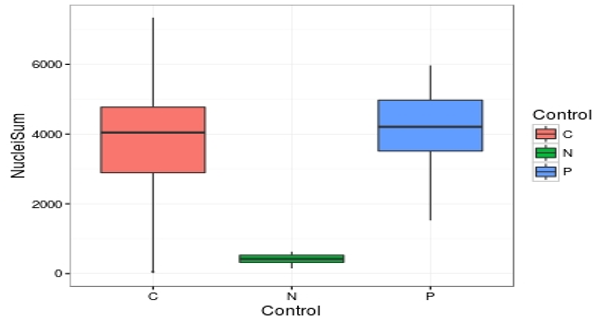
Figure 3. Pilot screen cell viability results. After 4 days incubation with compounds, the cells were fixed and stained with DRAQ5 as described.
Images were acquired (4 slices, 4 fields per slice in each well) and analyzed using CellProfiler. There is good separation between the positive (P) and negative (N) controls, with the compound (C) results spread between.
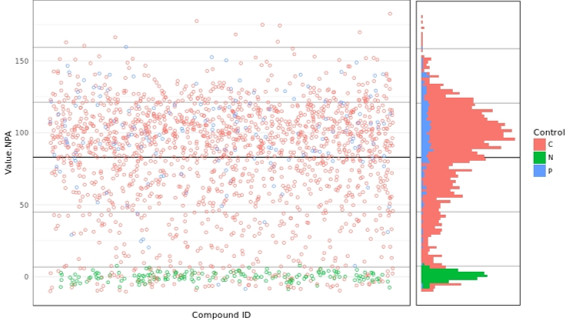
Figure 4. Scatterplot presentation of hits. The scatterplot of the results shows a fairly high hit rate at the compound concentration used (20 µM).
Conclusions
Better cell based assays are needed for robust and reproducible drug screening in a HTS format. In this study we have developed a reliable HepG2 liver cell 3D micro-tissue assay using GrowDex in an HTS-capable 384 well format. In the process, we have simplified the assay workflow and carried out a small pilot screen to demonstrate the utility of the assay. The results show the applicability of GrowDex as a matrix for 3D cell culture based drug screening in HTS and drug discovery.