Joanne Ossai1, Jonathan Sheard1,2 and Darius Widera1
1Stem Cell Biology and Regenerative Medicine Group, University of Reading, 2Sheard BioTech Ltd
Introduction
Glioblastoma multiforme (GBM) is the most common and aggressive brain tumour. Cell culture plays an indispensable role in the cancer drug discovery process. Notably, the majority of in vitro testing of anti-cancer drugs is carried out on traditional two-dimensional (2D) cell cultures. However, 2D cell culture only poorly represents the three-dimensional (3D) tumour microenvironment. Moreover, the high stiffness of plastic and glass substrates commonly used for 2D culture exceeds that of brain tissue resulting in stretched and unnatural morphology. The inability of 2D monolayers to closely approximate the pathophysiological conditions, has been shown to alter properties such as cell proliferation, morphology, gene and protein expression and ultimately, drug responses. Cultivation as self-assembled spherical clusters known as spheroids represents a widely used three-dimensional cultivation method for glioma cells. However, due to the compact nature of these `gliomaspheres`, the diffusion of nutrients and growth factors can be compromised [1, 2]. This highlights the need for novel 3D cell culture models that could act as better disease models and more sophisticated predictors in drug development [3].
Here, we developed a novel method for the 3D cultivation of the human glioblastoma cell line U251 within the nanofibrillar cellulose hydrogel, GrowDex®. We demonstrate that 3D cultivation of U251 cells in GrowDex results in an increased cell viability. Moreover, we show that in contrast to 2D, U251 cells in 3D respond in a linear manner to temozolomide which is an alkylating agent licensed for the treatment of newly diagnosed glioblastoma. This is in line with recent reports demonstrating that three dimensional matrices can affect cancer cell response to toxins [4].
Materials
- Human glioblastoma cell line U251-MG (ECACC 09063001, European Collection of Authenticated Cell Cultures)
- Complete Media: Dulbecco's Modified Eagle Medium: Nutrient Mixture F-12 (DMEM/F12) supplemented with 20% FBS, 1% Pen/Strep and 1% L-Glutamine (all Sigma Aldrich)
- GrowDex, 1.5% (Cat# 100 103 005, UPM)
- 3D culture: Low-attachment 96-well plates, (Cat# 83.3924.500,TC Plate 96 Well Suspension F, Sarstedt)
- 2D controls: Standard 96-well plates, (Cat# 83.3924.005 TC Plate 96 Well Standard F, Sarstedt)
- Cell proliferation kit II XTT (Sigma Aldrich)
- 4% Paraformaldehyde (PFA, Sigma Aldrich)
- 4′,6-Diamidine-2′-phenylindole dihydrochloride (DAPI, Sigma Aldrich)
- Phalloidin Atto 550 (Sigma Aldrich)
- Centrifuge 5810 R, Eppendorf with a rotor suitable for 96 well tissue culture plates
- Temozolomide (Sigma Aldrich)
- Spectra Max 340PC384 ROM v1.21 plate reader (Molecular Devices, Wokingham, UK)
- EVOS Live imaging system (ThermoFisher)
- Nikon A1 plus Confocal Microscope System (Nikon Instruments Europe B.V. United Kingdom)
Method
- U251 cells in 2D and 3D were cultured in complete media and incubated in a humidified incubator at 37°C and 10% CO2.
- For 2D culture, cells were resuspended in complete media at a concentration of 1000 cells/μL and cultivated in 96 well cell culture plates.
- For 3D culture, cells were mixed with complete media and GrowDex to provide a final concentration of 100μL of 0.1, 0.2 or 0.4% GrowDex seeded with 1000 cells/µl. Embedded cells were transferred into 96-well tissue culture plates and fed with 50µl complete media. Cells were maintained in a humidified incubator at 37°C and 10% CO2 and the medium was replaced every 2 to 3 days.
- For the viability assessment, 50μL the of XTT labelling mixture, was added to each well of the 96 well plate and the plates were incubated for 4 hrs in a humidified incubator at 37°C, 10% CO2 followed by measurement of the absorbance of the XTT formazan metabolite.
- For the phalloidin/DAPI staining, the culture media was carefully removed from cells in 96-well plate. Cell cultures were fixed with 4% (w/v) paraformaldehyde solution in phosphate-buffered saline (PBS) for 20 minutes. The fixative solution was removed, and cells were washed with PBS. Cells were permeabilized with 0.1% (v/v) TritonX-100 in PBS for 30 minutes at room temperature. To remove the permeabilization solution, the plate was centrifuged at 100g for 30 seconds. Lyophilisate of phalloidin (10mnol) was dissolved in 500µl of methanol. The cells were stained overnight at 4°C using 30µL of the phalloidin stock diluted in 1ml PBS containing DAPI (1:1500).
- Phase contrast images were taken using the EVOS imaging system whereas the phalloidin and DAPI stained cells were imaged on the Nikon A1 plus Confocal Microscope System
- U251 cells cultivated in 2D and 3D were exposed to increasing concentrations of temozolomide at 37°C at 10% CO2 for 48 h and subjected to XTT-based viability assessment.
Results
Phase contrast microscopy and XTT viability assays revealed that U251 grow homogenously within GrowDex and that 3D cultivation results in a significantly increased cellular viability compared to 2D controls (Figure 1). Moreover, we show using confocal laser scanning microscopy that U251 cells are evenly distributed in all three dimensions of the hydrogel (Figure 2). Finally, a drug response assay showed that 3D cultivation leads to a linear reduction of viability in response to the cytotoxic temozolomide stimulus (Figure 3).
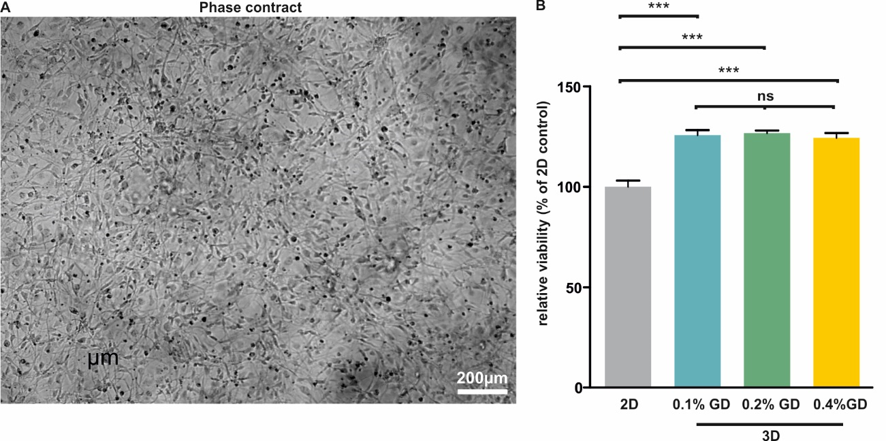
Figure 1. GrowDex supports 3D cultivation of U251 glioblastoma cells. A) U251 cultivated in 3D within 0.2% GrowDex form homogenous networks and display normal morphology. Scale bar: 200μm. B) XTT assay revealed that U251 cells cultivated within 0.1, 0.2, and 0.4% GrowDex show significantly increased cellular viability compared to the 2D control. Data is displayed as mean of three independent experiments ± SEM. ns: not significant. ****P <0.001.
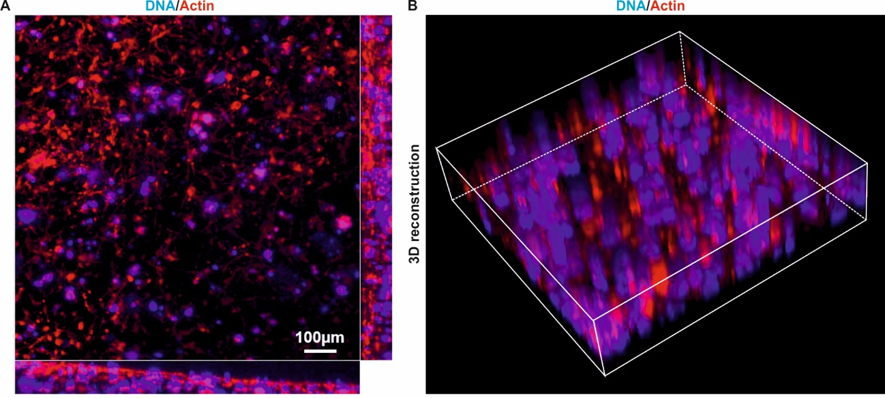
Figure 2. U251 cells are evenly distributed in all three dimensions of the GrowDex. Cells were cultivated in 0.2% GrowDex and stained for actin with Atto-coupled phalloidin. Nuclear counterstaining was performed with DAPI. A) Actin staining and confocal laser scanning microscopy revealed an isotropic distribution of cells in all three dimensions of the GrowDex hydrogel. Scale bar: 100μm. B) 3D reconstruction.
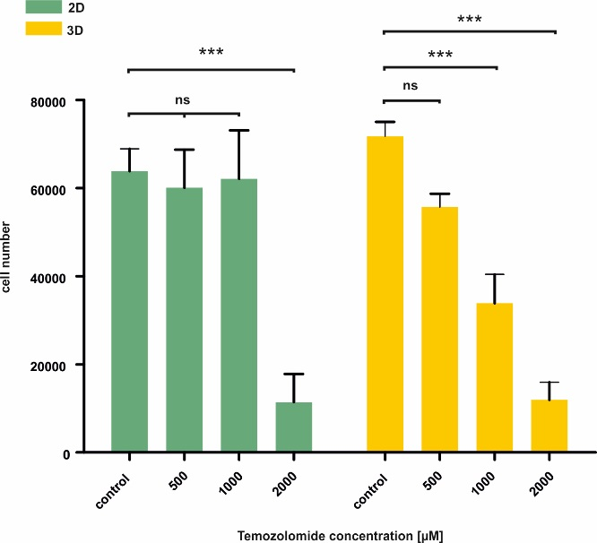
Figure 3. 3D cultivation within GrowDex results in a linear response to the cytotoxic drug temozolomide. U251 cells cultivated in 2D and 3D were exposed to increasing concentrations of temozolomide followed by assessment of viability using XTT. Statistical evaluation of the data revealed that cells within 3D respond to the toxic stimulus in a linear manner whereas only high concentration of temozolomide significantly reduce the viability when cultured in 2D. Data is displayed as mean of three independent experiments ± SEM. ns: not significant. ****P < 0.001.
Conclusions
Our results show that GrowDex is a suitable scaffold for 3D cultivation of the human glioblastoma cell line U251. Moreover, we show that GrowDex leads to an increased viability over concentration range from 0.1-0.4% when compared to 2D culture. Finally, we demonstrate that 3D culture in GrowDex can mediate a more linear drug response in comparison to conventional 2D culture. The results indicate that 3D cultivation of glioma cells within GrowDex might better reflect the pathophysiological environment and could improve current 2D cell-based drug screening platforms. Notably, our method could be easily adopted for primary glial cells, other glioma cell lines, and primary glioma cell cultures.
References
- Edmondson, R., et al. (2014). "Three-Dimensional Cell Culture Systems and Their Applications in Drug Discovery and Cell-Based Biosensors." Assay and Drug Development Technologies 12(4): 207-218.
- Laks, D. R., et al. (2016). "Large-scale assessment of the gliomasphere model system." Neuro Oncol 18(10): 1367-1378.
- Matak, D., et al. (2017). "Colony, hanging drop, and methylcellulose three dimensional hypoxic growth optimization of renal cell carcinoma cell lines." Cytotechnology 69(4): 565-578.
- Zustiak, S. P., et al. (2016). "Three-dimensional matrix stiffness and adhesive ligands affect cancer cell response to toxins." Biotechnol Bioeng 113(2): 443-452.