Jonathan Sheard, Ph.D., Sheard BioTech Ltd & University of Reading, UK
INTRODUCTION
A robust and reproducible cell viability assay is an essential tool for evaluating changes in cell health and behaviour, for example in drug efficacy studies [1]. There are several ways to measure cell viability, such as dye exclusion [2], direct DNA stains and dyes, as well as the detection and quantification of fluorescence or luminescence by enzymatic activity [3]. Additionally, it is known that cells grown in conventional two-dimensional (2D) monolayer cultures may lose many tissue-specific properties, protein expression profiles as well as changes to drug sensitivity [4, 5].
Therefore, in order to obtain better in vivo relevancy, researchers have turned their attention to the development of 3D cell-based assays where in vivo-like properties are retained, offering a more relevant model [6, 7]. Fibroblasts play a crucial part in wound healing [8], formation of extracellular matrix and inflammation [9], as well as cancer [10]. Thus, analysis of normal and disrupted fibroblast behaviour and viability are important in further understanding these roles.
In this proof of concept study, we investigated the use of the Promega CellTiter-Glo® (CTG), CellTiter-Glo 3D® (CTG-3D) and RealTime-Glo™ (RTG) luminescent assays to measure the presence and production of ATP from viable cells. The cell model used, was a 3D assay using human dermal fibroblasts which were cultured in GrowDex® hydrogel.
GrowDex is an animal free hydrogel produced from birch wood-derived nanofibrillar cellulose. Ambient handling and the shear thinning properties of GrowDex make it a ready to use product which and can be mixed directly with cells and culture media, without the need for cross-linking steps, allowing the hydrogel to be customized to provide a controlled and completely animal free cell environment [11]. Additionally, due to the natural material characteristics of the hydrogel small molecules, such as those included in the Promega viability assays, can easily diffuse through the gel to gain access to the cells.
Both CTG and CTG-3D are endpoint lytic assays. Beetle Luciferin, in combination with ATP (from the cells) and O2, is enzymatically transformed by the Ultra-Glo™ Recombinant Luciferase to produce light. The luminescent signal generated as a result of the enzymatic activity can be directly correlated to the concentration of ATP present in the cell lysate samples. The RTG viability assay offers an alternative non-lytic bioluminescent method for the measurement of cell viability in real time by measuring the reducing potential of cells and thus metabolism. A cell permeant pro-substrate (MT Cell Viability substrate) is added to cells, which is then metabolically reduced by viable cells to produce the NanoLuc® substrate. The
NanoLuc substrate diffuses from cells into the culture medium where the NanoLuc enzyme acts on the substrate to produce a luminescent signal.
In this study, the performance of all three Promega cell viability kits are evaluated in 3D cellbased assays with human dermal fibroblasts.
MATERIALS
- Normal human dermal fibroblasts – Neonatal (NHDF, Cat No. CC-2509, Lonza)
- Complete media: DMEM-High Glucose (Cat No. D5671-500ML, Sigma) supplemented with 20% heat inactivated foetal bovine serum (Cat No. F9665, Lot: RNBG8272, Sigma), 1% L-glutamine (200 mM, Cat No. G7513-100ML, Sigma), 1% penicillin with streptomycin (10,000 units penicillin and 10 mg streptomycin per ml, Sigma) and 5 ng/ml fibroblast growth factor (FGF2, Cat No. AF-100-18B, Peprotech).
- GrowDex® 1.5% (Cat No. 100 103 005, UPM Biomedicals)
- CellTiter-Glo® (CTG, Cat No. G7571, Promega)
- CellTiter-Glo® 3D (CTG-3D, Cat No. G9682, Promega)
- RealTime-Glo™ (RTG, Cat No. G9712, Promega)
- Triton X-100 (Cat No. T8787-50ML, Sigma)
- Black clear bottom 96 well plate (Cat No. 655090, Greiner Bio-One)
- SpectraMax iD3 Microplate Reader (Molecular Devices)
METHODS
1. NHDFs were culture expanded in 2D in complete media and incubated at 37°C with 10% CO2. Following trypsinization, cells were resuspended in complete media at a concentration of 1x106 cells/ml.
2. Stock GrowDex (1.5% w/v) was diluted, firstly with culture media and then with media containing NHDFs, to make a final working solution with a GrowDex concentration of 0.4% and 1x106 cells/ml [12].
3. Working example: for 1 ml of final working solution, add 266 μl of GrowDex to 634 μl of complete media and mix thoroughly whilst avoiding bubbles. Then add 100 μl of cells which are in suspension at a stock concentration of 1x106 cells/ml and mix thoroughly. This will give you a 1ml working solution with 1x105 cells/ml in 0.4% GrowDex.
4. Subsequently, 50 μl of cells embedded in 0.4% GrowDex was pipetted into each well of a 96 well plate.
3. A ’no cell control’ made up of 0.4% GrowDex (without cells) in complete media was prepared and 50 μl pipetted into wells in the same plate, to measure any interaction and effects of the hydrogel on luminescent measurement or signal intensity.
4. After 30 mins, 50 μl of complete media was carefully added to all wells on top of the 0.4% GrowDex layer, by pipetting against the inside edge of the well.
5. All cultures were incubated at 37°C with 10% CO2 in a humidified incubator for 24 h and then the viability assays were performed.
6. Triton-X 100 was used as a cell toxin at a final concentration of 0.01% since it permeabilizes the membrane of cells, allowing ATP to leak out leading to cell death. 10 μl of complete media with or without Triton-X 100 was added 5-10 min prior to the addition of the viability kit reagents.
7. Reagents for the CTG, CTG-3D and RTG were prepared according to manufacturers instructions. 110 μl of each reagent was added (equal 1:1 of total well volume) to their respective wells. Following addition of the reagents to the wells, luminescence measurements were taken after 30 min for the CTG and CTG-3D viability kits. RTG assay luminescence measurements were taken at 30 min and then hourly from 1 h
through to 4 h after addition of the reagents.
RESULTS
Viability of NHDFs cultured in 0.4% GrowDex was successfully measured using all three of the luminescence assay kits provided by Promega. The CTG and CTG-3D assays (Fig. 1A) both showed a clear difference between untreated (green) and Triton X-100 treated (purple) cells 30 min after addition of the toxin and detection reagent. Additionally, the CTG-3D assay proved to be more sensitive as a much brighter luminescent signal was detected.
The RTG viability assay measuring NHDF viability in real time for the duration of the 4 h experiment was also successful. As shown in Fig. 1B, untreated cells (green) remain viable with an increased production of luminescent signal throughout the study. Permeation of the cell membrane by Triton X-100 (purple) showed a reduction in luminescent signal at 30 minutes and total loss of signal observed by the 2 hour time point.
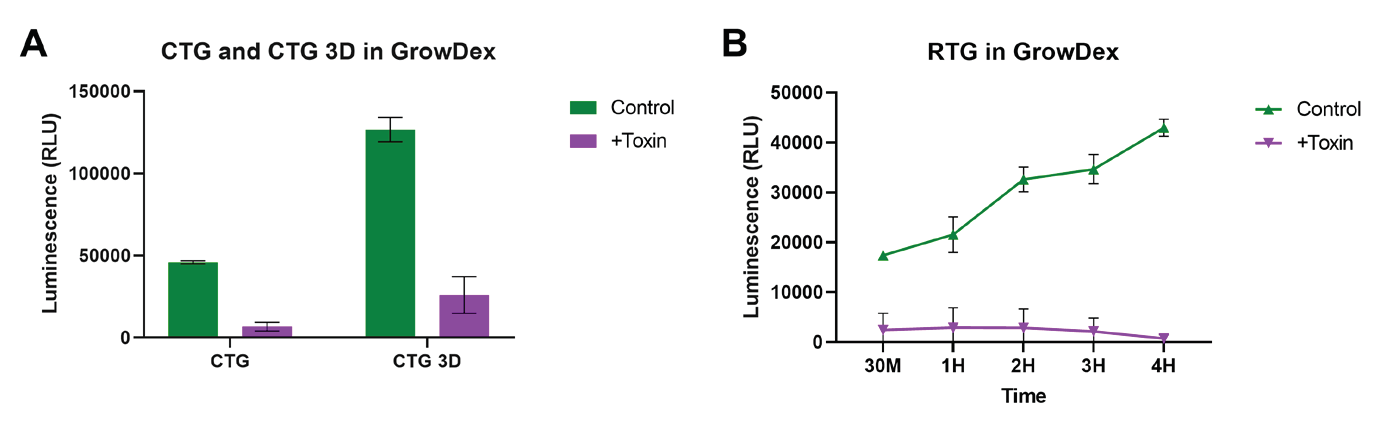
Figure 1. Cell viability was assessed using the CellTiter-Glo, CellTiter-Glo 3D and RealTime-Glo viability kits. NHDF (1x105 cells/ml) were embedded and cultured in 0.4% GrowDex for 24 h. Cells were treated with and without 0.01% Triton-X 100 (Toxin) 5-10 mins prior to measuring viability. Cell viability was measured using the lytic viability kits, CellTiter-Glo and CellTiter-Glo 3D (A) 30 min after treatment with toxin. The nonlytic real-time assessment of cell viability was measured using the RealTime-Glo viability kit (B) at 30 min then at 1 h intervals up to 4 h after treatment with toxin. Data shown as mean ± SEM
from N=2-3 experiments.
CONCLUSIONS
The lytic (CTG and CTG-3D) and non-lytic (RTG) approaches for the measurement of cell viability based on quantification of ATP from metabolically active cells are commonly used and reliable techniques. Here we show that the Promega viability kits, CellTiter-Glo, CellTiter-Glo 3D and RealTime-Glo luminescent cell viability assays can be successfully employed for the measurement of NHDFs cultured in a GrowDex 3D cell-based model. Measurements can be either endpoint (CTG and CTG-3D) or real time continuous (RTG) read-out assays depending on the researcher’s requirements. Additionally, the greater sensitivity of CTG-3D allows for the detection of low levels of cell viability within more complex models. Across the 4 h viability assessment the RTG assay kit, the toxin was seen to reduce signal intensity to minimal levels of luminescence detection, however untreated cells continued to increase luminescence signal, indicating a metabolically active and therefore viable population.
Here, we have successfully shown the culture and viability assessment of human dermal fibroblasts within GrowDex. Due to the unique material characteristics of GrowDex molecules, such as those provided in the Promega kits, can diffuse easily throughout the gel. GrowDex hydrogels with Promega cell viability kits enable fast and reproducible 3D cell culture assays for accurately measuring cell viability. There is no interference of the luminescent signal by GrowDex which means the combination of Promega cell viability assays and GrowDex hydrogel make a perfect combination for any 3D cell-based study.
REFERENCES
1. Huber, J.M., et al. (2016). "Evaluation of assays for drug efficacy in a threedimensional model of the lung." Journal of Cancer Research and Clinical Oncology 142(9): p. 1955-1966.
2. Strober, W. (1997). "Trypan Blue Exclusion Test of Cell Viability." Current Protocols in Immunology 21(1): p. A.3B.1-A.3B.2.
3. Ramirez, C.N., et al. (2010). "Cell viability assessment: toward content-rich platforms." Expert Opinion on Drug Discovery 5(3): p. 223-233.
4. Melissaridou, S., et al. (2019). "The effect of 2D and 3D cell cultures on treatment response, EMT profile and stem cell features in head and neck cancer." Cancer Cell International 19(1): p. 16.
5. Ossai, J., et al. "Human glioblastoma (U251): 3D model demonstrates increased cell viability and linear drug response over 2D." GrowDex Application Note 25.
6. Sheard, J.J., et al. (2019). "Optically Transparent Anionic Nanofibrillar Cellulose Is Cytocompatible with Human Adipose Tissue-Derived Stem Cells and Allows Simple Imaging in 3D." Stem Cells International 2019: p. 12.
7. Azoidis, I., et al. (2017). "Three-dimensional cell culture of human mesenchymal stem cells in nanofibrillar cellulose hydrogels." MRS Communications 7(3): p. 458-465.
8. Bainbridge P.1 , B.B.C. (2013). "Wound healing and the role of fibroblasts." Journal of Wound Care 22(8): p. 407-412.
9. Lo, D.D., et al. (2015). Chapter 51 - Differences in Foetal, Adult Skin and Mucosal Repair. Stem Cell Biology and Tissue Engineering in Dental Sciences. A. Vishwakarma, et al. Boston, Academic Press: 691-702.
10. Kalluri, R. and Zeisberg, M. (2006). "Fibroblasts in cancer." Nature Reviews Cancer 6(5): p. 392-401.
11. Coots, K.S., et al. "Neuronal Differentiation of Human Embryonic Stem Cell Derived Otic Neural Progenitors Using a Nanofibrillar Cellulose and Hydrogel Scaffold and Polyhedrin Delivery System." GrowDex-T Application Note 5.
12. UPM-Biomedicals "Recommended Procedure For Diluting and Mixing GrowDex®-T." GrowDex-T Application Note 1.
You can order products online at: www.upmbiomedicals.com/store
Or contact us at biomedicals.sales@upm.com for a quotation or to place an order.